What is Bayesian Linear Regression? (Part 2)
Matthias Werner
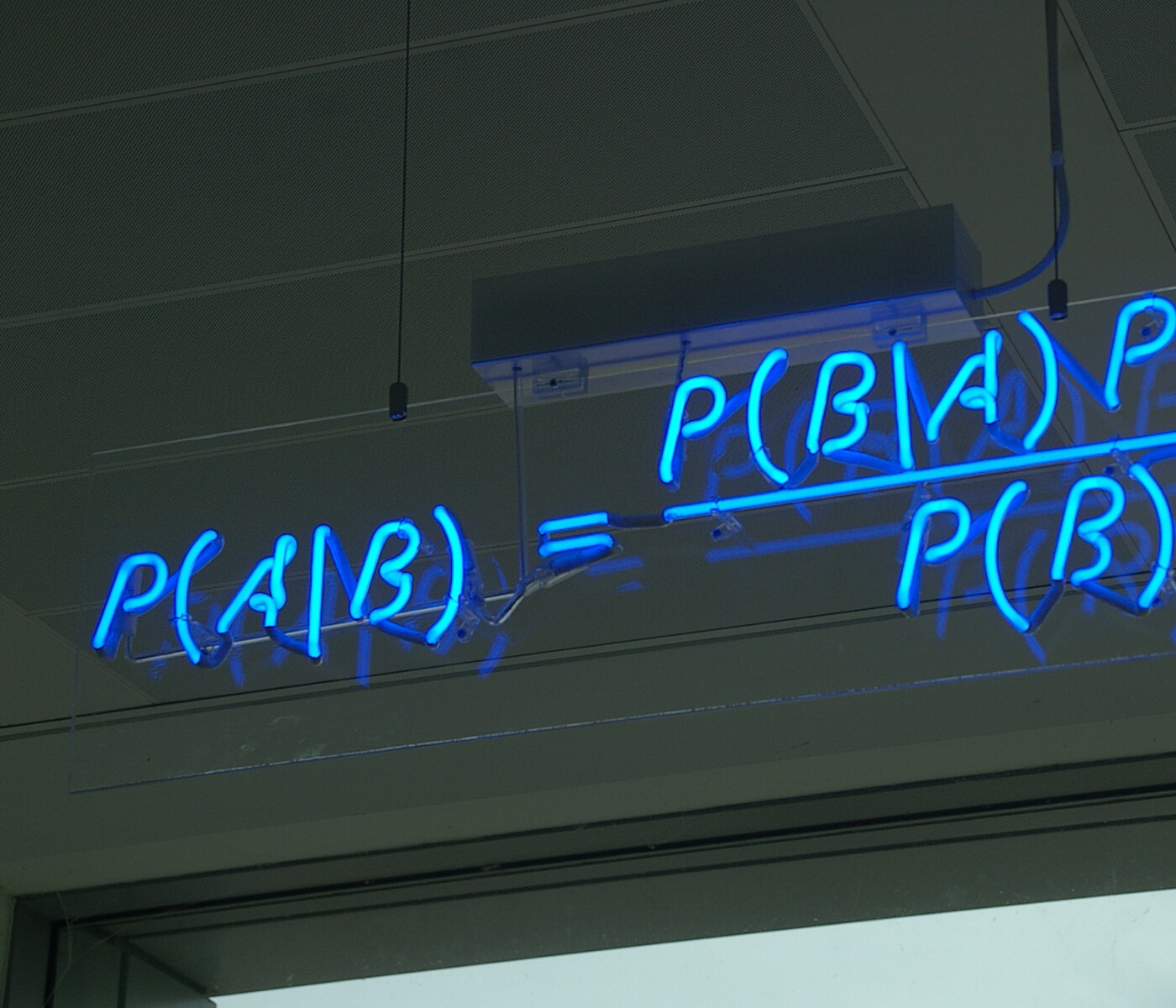
In my previous blog post I have started to explain how Bayesian Linear Regression works.
So far, I have introduced Bayes' Theorem, the Maximum Likelihood Estimator (MLE) and Maximum A-Posteriori (MAP). Now we will delve into the mathematical depths of the details behind Bayesian Linear Regression.
Bayesian Linear Regression
We have used Bayes' theorem to justify estimating the model parameters with regularization, but we are still using point estimates of the model parameters $$\vec{w}$$. Bayes' theorem could theoretically give us access not just to the maximum of the posterior distribution as in MAP, but allow us to use the distribution $$p(\vec{w} | D)$$ itself. This will allow us to estimate the standard deviation of our prediction. We will use conjugate priors to compute $$p(\vec{w} | D)$$. Conjugate priors are prior distributions $$p(\vec{w})$$ which are in the same family of distributions as the posterior $$p(\vec{w} | D)$$. As in the examples above, we will assume the likelihood $$p(y_i | \vec{x}i, \vec{w})$$ and the prior $$p(\vec{w})$$ to be Gaussians. A Gaussian times a Gaussian is still a Gaussian. As it can be seen in Bayes' Theorem, the formula for normalization and Bayes' Theorem, the prior of the training data $$p(D)$$ is essentially a normalization constant and will not influence the general shape of $$p(\vec{w} | D)$$. So we already know $$p(\vec{w} | D)$$ will be a Gaussian. For a Gaussian we only need to figure out the mean vector $$\vec{\mu}_w$$ and the covariance matrix $$\Sigma_w$$ of $$p(\vec{w} | D)$$ and then can infer the normalization from there.
Let's do the math!
So how to start? Let's first write out the distribution we are looking for. We will then make a reasonable choice of $$\vec{\mu}_w$$ and $$\Sigma_w$$. Using Bayes' Theorem and this choice we show that we recover the posterior distribution $$p(\vec{w} | D)$$.
As just discussed above, assuming the likelihood $$p(yi|\vec{x}i, \vec{w})$$ to be Gaussian and using a conjugate Gaussian prior we know the posterior to look something like this (in order to refer to the equation more easily we call it GOAL):
The proportionality $$\propto$$ is due to the fact that we will take care of normalizing the distribution at last, as discussed in the section for the "Bayesian" workflow. Exploiting the proportionality we can use the numerator of Bayes' Theorem to obtain
Now let's try to rewrite the right-hand side into this shape. Given that the likelihood $$p(y_i | \vec{x}_i \vec{w})$$ and the prior $$p(\vec{w})$$ are Gaussians, we can write (equation START)
In order to write more comfortably, we introduce vector and matrix notation for our data points. Consider the following matrix of data points $$X$$ such that the columns are our data points $$\vec{x}_i$$, i.e.
and the row vector
So we simply defined a matrix $$X$$ where the columns are the x-values for the regression problem and a vector $$\vec{y}$$ where the elements correspond to the respective y-values. Let's re-write the likelihood term on the right-hand side a little bit:
where $$I$$ is the identity matrix.
In order to save ourselves some trouble with the notation, we can already identify the mean vector and covariance matrix of the posterior distribution (equation MOMENTS)
Notice how we only need the training data $$X$$ and $$\vec{y}$$ and some assumption about the noise $$\sigma_{\epsilon}^2$$ to compute the MOMENTS. This might appear a little out of the blue, but we will explain now why it is reasonable to choose this. Consider the shape of the exponent in GOAL (equation QUADRATIC):
This equation can be easily shown to hold when considering that $$\Sigma_w^{-1}$$ is symmetrical. Using MOMENTS, QUADRATIC and some patience in INTERMEDIATE we get
Here we notice that $$\exp \left[ -\frac{\vec{y} \vec{y}^T} {2\sigma_{\epsilon}^2} + \frac{1}{2} \vec{\mu}_w^T \Sigma_w^{-1} \vec{\mu}_w \right]$$ is constant with respect to $$\vec{w}$$. This means we can take it as a factor and make it disappear into a proportionality:
We find, using the definitions MOMENTS we recover the posterior distribution GOAL of the model parameters up to normalization. The normalization of a multi-variate Gaussian is known once the covariance matrix $$\Sigma_w$$ is known, but in the example we looked at in the first part of this article, we did not need to explicitly compute the normalization. The mean vector and covariance matrix in MOMENTS can be calculated from the training data $$D$$ and then allow us to make predictions with uncertainty estimation, as shown in the previous article.
To summarize
We learned how to use Bayes' Theorem to various degrees:
We ignored Bayes and ended up with a simple MLE. MLE for linear regression turns out to be identical to minimizing the sum of squared errors.
We used Bayes' Theorem for a point estimate and got MAP. MAP for linear regression and a Gaussian prior of the parameters turns out to be equivalent to MLE with L2-regularization.
Last but not least, we used Bayes' Theorem to compute the entire posterior distribution of the model parameters using conjugate priors. Once we know the mean vector and the covariance matrix of the posterior, we can analytically compute the mean prediction and the standard deviation / variance of this prediction. Furthermore, we can sample individual sets of parameters.
I hope this article gave an in-depth introduction to BLR and how it relates to various popular methods of model parameter estimation. If you have notes or question do not hesitate to contact us!
